Dr Clare Sansom, Senior Associate Lecturer in Birkbeck’s Department of Biological Sciences reports on Dr Salvador Tomas’ talk, which explored the hypothesis of abiogenesis, which assumes life arose spontaneously from non-living matter, a few billion years ago. The talk focused on research directed to the development of protocells, which are produced in the laboratory as plausible ancestors of living cells, and can be used as models to study abiogenesis. In the future, scientists may be able to use this knowledge to create programmable, cell-like robots.

Tekno the Robotic Puppy, credit: Toyloverz
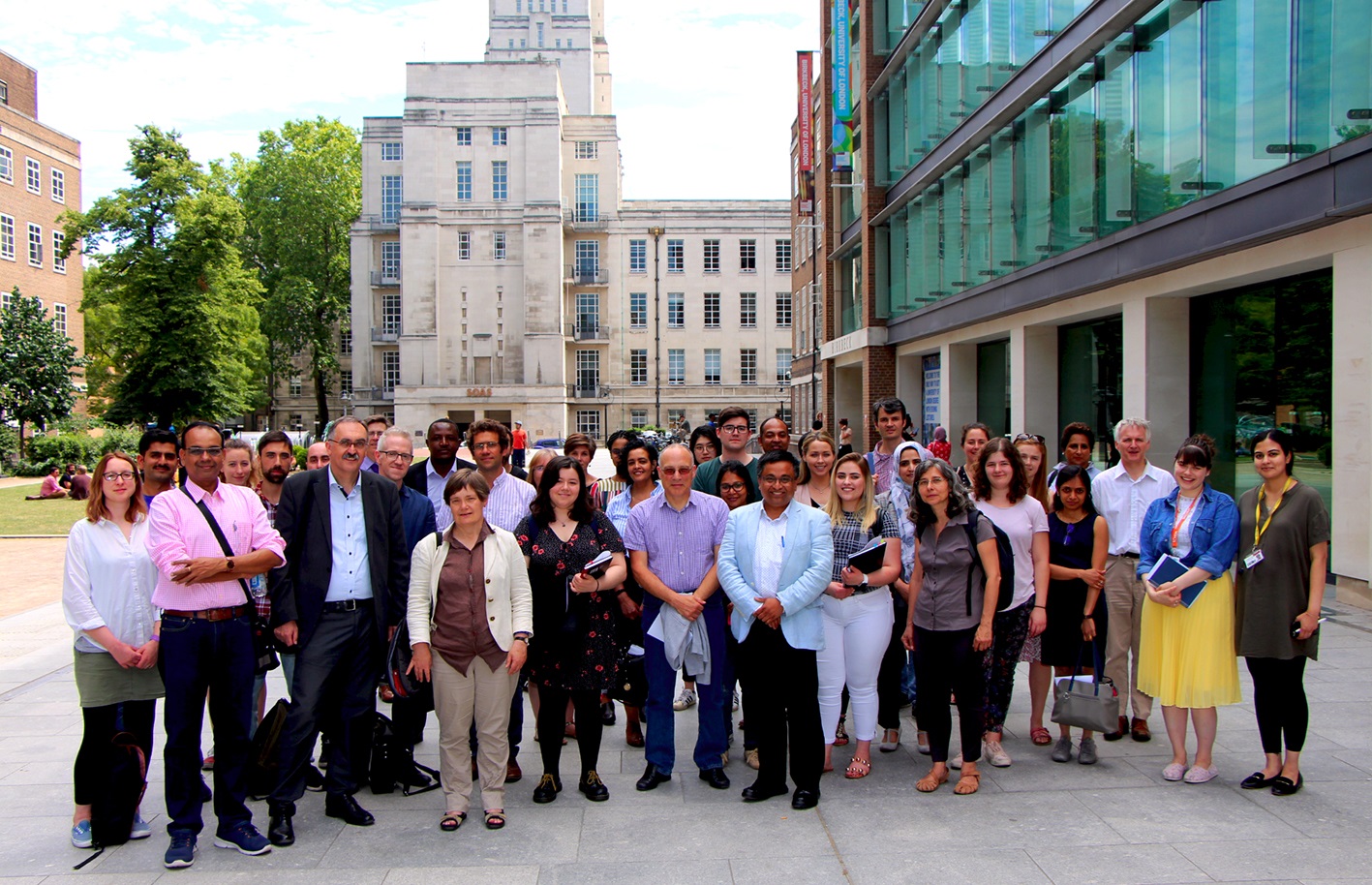
Birkbeck Science Week 2019 kicked off with a talk by Dr Salvador Tomas of the Department of Biological Sciences, intriguingly titled ‘Synthesising Life’. Introducing Salvador, the Executive Dean of the School of Science at Birkbeck, Nicholas Keep, explained that he had taken both his degrees at the Universitat de les Illes Balears in his native Balearic Islands. He moved to Birkbeck to set up his lab in 2006 following postdoctoral study in Sheffield, and he now holds the position of Senior Lecturer in Chemical Biology.
His lecture was every bit as engaging as its title suggests. He started by asking the question ‘what is life?’ and illustrated the answer by comparing a ‘cyberdog’ with the common-or-garden variety. At a basic level, both dog and cyberdog can be thought of as a network of transistors or cells that responds to input signals in different ways, but while the cyberdog is programmed to carry out whatever (presumably) menial tasks its owner demands, the dog is programmed for survival. This led to a formalised definition of ‘life’, as ‘a self-sustained chemical system capable of undergoing Darwinian evolution’. Furthermore, if you zoom in hundreds of millions of times, the dog’s equivalent of the cyberdog’s uniform network of transistors is the bewildering complexity of ‘molecular machines’ inside every living cell.
The question of ‘how life came to be’ is perhaps almost as old as humanity itself. A few centuries ago speculations centred on the idea of ‘spontaneous generation’, suggesting that (for example) fish might have arisen directly from water or mice from hay. The development of pasteurisation in the mid-nineteenth century helped disprove this theory. We now understand that all living (and extinct) organisms evolved from a simple organism known as LUCA – short for the Last Universal Common Ancestor – but this begs the question: where did LUCA come from? To answer this question, you need to go back to the kind of conditions that scientists believe to have existed on an early Earth: a chemically rich ‘warm puddle’ of liquid in an oxygen-poor environment, much like those found in underwater volcanoes today.
LUCA would have been a single-celled organism containing a minimum set of biomolecules necessary for life, all coded for by a minimal segment of DNA, and, self-evidently, all its precursors must have been non-living. For decades, scientists have been trying to recreate the process of ‘abiogenesis’ by providing simple molecules with energy in a similar environment and investigating whether more complex molecules, the ‘building blocks’ for LUCA’s DNA and proteins, might be able to form. So far, it has proved possible to make the basic building blocks of proteins, the amino acids, and even, in some circumstances, to join several amino acids into a short chain, but not to connect hundreds of them to form a complete protein. Nucleic acids, the building blocks of DNA, are proving even more intractable.
Building blocks become biomolecules through a process in which each two units – amino acids or nucleotides – are joined together with the loss of a water molecule. This process requires energy, but the opposite one, in which the bond between the units is broken, can be spontaneous. Salvador used a set of simple blocks to illustrate how populations change over time, as combinations such as ‘AB’ are ‘born’ and ‘die’. If AB, for example, is made ‘sticky’ so it attracts more copies of A and B, it becomes ‘autocatalytic’ (that is, it helps form itself) and the AB population burgeons. Or at least, it does so until A or B is depleted, when an ‘extinction event’ occurs. The system becomes more complex with the addition of an energy supply and further building blocks, and it becomes possible to see how collections of units with specialist functions could evolve. Some types would specialise in storing information (the ancestors of DNA) and others in promoting bond formation (the ancestors of proteins).
This would be a resourceful molecular system, capable of building its own building blocks, but it would have one major disadvantage: its survival would depend on the proximity of different types of molecule. If it were in the ‘warm puddle’ of the early Earth, a single rainstorm could blow it away. Keeping the components together requires a third type of biomolecule. Lipids are molecules with a long ‘water-hating’ tail and a short ‘water-loving’ head, and in water they form double layers with the tails pointing towards each other. These lipid bilayers often form spherical vesicles, and any primitive biomolecules trapped inside such a vesicle will stay together come what may.
Vesicles containing both ‘DNA-like’ and ‘protein-like’ molecules can be thought of as ‘protocells’: or, if you like, putative ancestors of the ancestors of LUCA. Salvador explained that his own contribution to the evolving story of synthesising life was in exploring the chemistry inside such protocells. Something like a protocell is almost certain to have existed, and this will have evolved to be better programmed for survival through developing more efficient chemical ways of making use of resources, storing and using energy, and responding to stimuli. Reproducing this process by adding molecular machines and efficient, specialist switches to a blank vesicle or protocell can generate cell-like robots. Initially, these are likely to have a variety of useful but quite mundane functions in, for example, targeted drug delivery, but eventually they might do more: ‘life, but not as we know it’, perhaps?
Salvador ended by asking two questions: can we synthesise life, and if so, should we? Most of his audience agreed with him that the first was ‘not done yet, but seems likely in the near future’. Interestingly, however, a majority thought that it might be too risky to take very far.